Feature Engineering
- Determines node's features from the domain (experts)
- Compute node's features from the graph structure
- Degree
- Centrality
- Betweenness
- Closeness
- DeepWalk, node2vec, ...
- Spectral embedding
Node Degree
Simply taking the degree of a node as a feature.
- Indicates how important is an user in a social network (influencer)
- Only very local information
- Too Limited
Node Centrality
Estimate the position of a node within the graph
- Closeness centrality
- Betweenness centrality
- Eigenvector centrality
Closeness Centrality
Measures how central a node is by calculating the reciprocal of the sum of the shortest path lengths from that node to all other nodes.
- In disconnected graphs, traditional closeness centrality isn't applicable. Alternative measures, like harmonic centrality, are used to handle infinite distances.
- Closeness centrality assumes information spreads along shortest paths, which may not always reflect real-world scenarios.
Betweenness Centrality
Quantifies the importance of a node within a network by measuring the number of shortest paths between pairs of nodes that pass through it. Nodes with high betweenness are critical connectors within the network.
Eigenvector Centrality
Measures a node's influence within a network by considering not only the number of its direct connections but also the importance of the nodes it connects to.
For a graph with adjacency matrix
-
-
-
Related to the PageRank algorithm
Spectral embedding
Laplacian EigenMaps
Use the eigenvectors of the Laplacian matrix as feature vectors
Example on KarateClub
Learn node embeddings
Learn representations instead of defining them explicitly
Shallow Encoding
Find a vectorial representation
Node Feature Matrix
Similarity Node Matrix
-
-
How to define
-
How to compute
DeepWalk
- Unsupervised learning
- Use Random Walks to explore the graph
- Transform node sequence to vectors using Word2Vec like approach
DeepWalk : Random Walks
- For each node, compute random walks
- Nodes composing a random walk constitute a sequence
Random Walk computation
- Transition Matrix
DeepWalk : Word2Vec for graphs
Learning node representations based on the similarity of their random walk distribution.
- Node sequences are sentences (as in NLP)
- Adaptation of Skip Gram model from word2vec to learn a node embedding.
- Nodes sharing random walks in the graphs are associated to similar vectors.
Node2vec
- Improvement over DeepWalk.
- Two new parameters to drive the random walks (BFS vs DFS)
- Allow to adapt the exploration to the graph
Node2vec
Tradeoff between local and global exploration
- p : probability to come back to previous node
- q :Favorize BFS or DFS
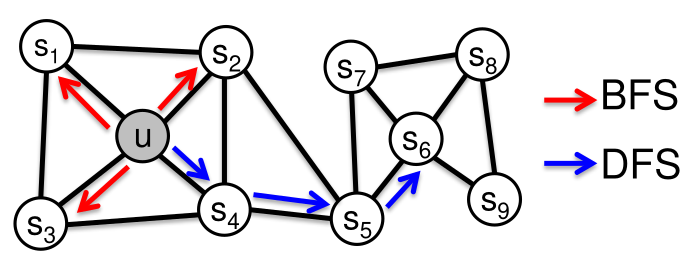
Example on KarateClub
Conclusion on node embeddings
- Useful for node level tasks
- Embedding can be defined a priori or learnt using the structure in a unsupervised way
- Widely used for network analysis
- Learning embeddings according to a downstream task : Next lesson !
Graph Embedding
- Associate a vector to the whole graph
- Permutation invriant function
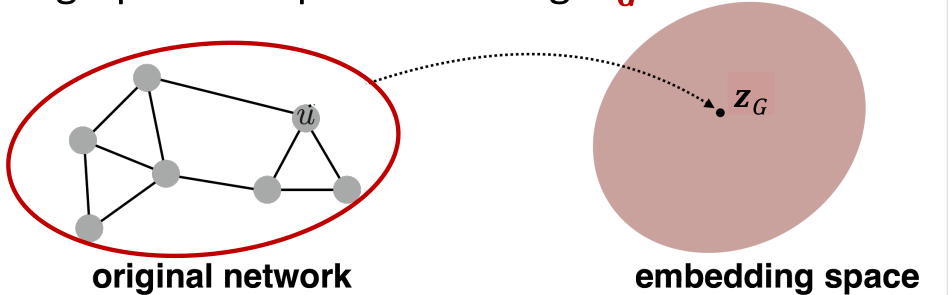
Simple Approach
Reach the invariance by an invariant aggregation function (sum, mean, etc)
Graph representation is a resume of the set of node representations.
Depends on the quality of node embeddings
Application Dependant Approach
Use a set of predefined graph features.
- Require an expert domain
- Sub Structures counting (more on that next)
- Fingerprints for molecular compounds (RDkit, OpenBabel)
- Molecular Descriptors (Avoid the graph problem)
Empirical Map
Consider that you are able to compute a distance
In your learning dataset, identify
- Depends on the quality (and number of anchors)
- Versatile approach
Topological embedding
Local similarities induce global similarity
- Define a set of substrutures
- Compute the number of occurences of each substructure in the graph
- Encode label information trough histograms [Sidere et al. 09]
- Considering more detailed labeled information may lead to huge vectors sizes
- Related to fingerprints and bags of patterns kernels
Global Approaches
Fuzzy multilevel graph embedding [Luqman et al.13]
Add basic global information like the number of nodes or edges in the graph.
Spectral embedding [Luo et al. 03, Caelli et Kosinov 04, Luo et al. 06]
Define an embedding from the spectrum of laplacian or adjacency matrix
Matrix Factorization
Find the best embedding to recreate a distance matrix
- Similarity matrix
- Factorize
Works only if d is an euclidean distance)
Graph Kernels
All seen embeddings approaches require to define explicitly the vector.
- Limits the expressivity
- The bigger the size, the more complex the computations
Kernel Theory
Kernel
- Symmetric:
- For any dataset
Reproducing Kernel Hilbert Spaces
-
Scalar product
-
Distance
-
A kernel is a similarity measure between objects.
Kernel Trick with SVM
Objective function
s.t
Decision function
Kernel Trick Example
Example
- No linear solution.
- Solution in
Graph Kernels
Graph Kernel Theory
- Graph kernel
- Similarity measure between graphs
- Use of natural representation of data (molecules)
- Implicit graph embedding in
- Access to kernel methods (SVM, KRR, etc.)
Kernel Definition
Graph kernel based on bags of patterns
- Extraction of a set of patterns
- Comparison between patterns
- Comparison between bags of patterns
Walks and Paths
- Patterns are walks or paths in the graph
- Walk: Sequence of nodes
- Path: Walk without repeated nodes
Tottering walk: Walk with a repeated node
Graphlets
Set of subgraphs having at most 5 nodes
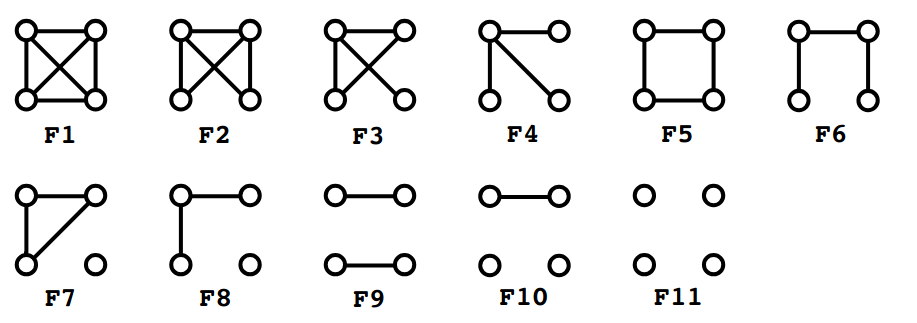
Treelets
Set of labeled sub-trees having at most 6 nodes
- Encode structural information
- Labeled patterns
- Limited set of structures
Kernel Definition
Counting function
Kernel definition
Graph similarity
Pattern Selection
- Different properties
- Domain Expert Approach: Selection of relevant descriptors for a given task
- Machine Learning Approach: Define the set of substructures according to an objective
Multiple Kernel Learning
Optimal combination of kernels:
or
- Each sub-kernel is weighted
- Selection of relevant treelets
- Weighting computed according to a given property
Weisfeler-Lehman Kernel
Principle : Aggregate neighboorhood information in an iterative fashion
- Related to WL Isomorphism Test
Question : How to derive an inexact similirity measure from this isomorphism algorithm ?
Weisfeiler-Lehman Kernel
- Count the number of common labels at iteration
Weisfeiler-Lehman Kernel
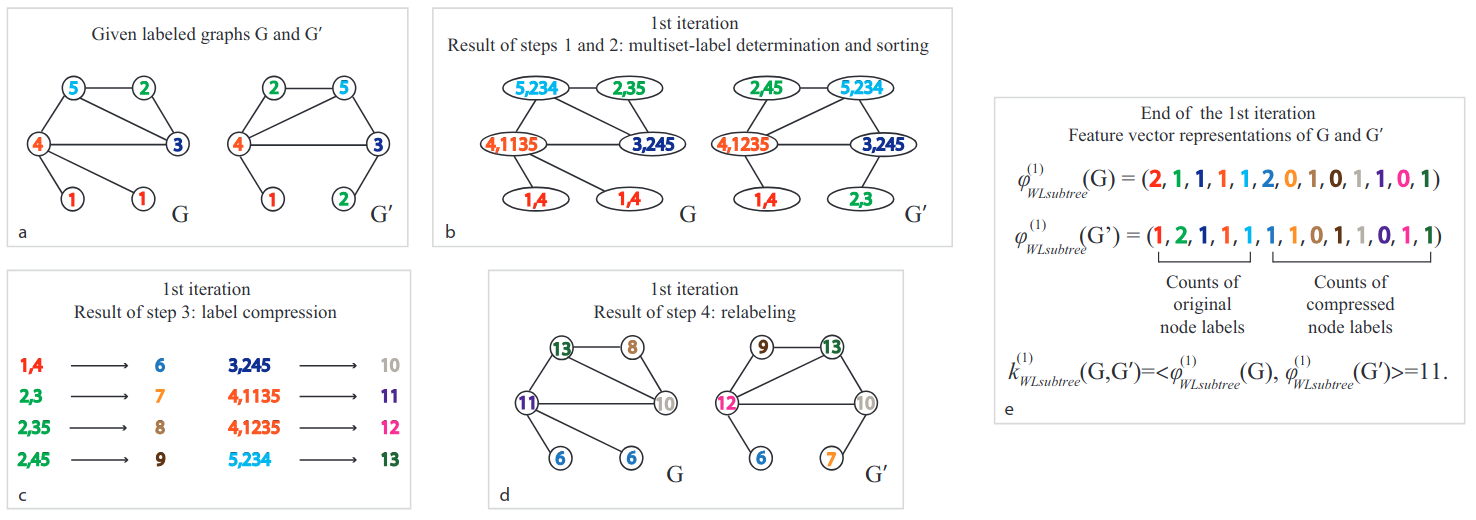
Weisfeiler-Lehman Kernel
- Efficient to compute
- Enrich the node description iteratively
- Widely used in many applications
- Fundations of modern Graph Neural Networks (next lesson)
Conclusion
- Node and Graph embeddings are essential for graph analysis
- Node embeddings are used for node level tasks
- Graph embeddings are used for graph level tasks
- Many methods exist to define embeddings
- Kernel methods are a powerful tool to define embeddings
- All embeddings are limited by the choices made